Reading our DNA is a new way of interpreting ourselves. Current trends suggest that one day, perhaps much sooner than we think, we will all have our own genomes sequenced. We may opt in for health reasons, curiosity about our ancestry, or to push the intellectual and emotional boundaries of what it means to be human. Some of us will just want to be part of the growing ‘genomics club’. It makes for interesting conversation around the water-cooler—or these days, the espresso machine.
There is increasing talk of sending all babies home from the hospital. This is just one step towards the use of genome information as a routine part of health care. How might babies with known genome sequences fare compared to those born without a crystal ball in their cradle? This is what Robert Green and Alan Beggs of Harvard University hope to find out in their BabySeq project, one of a new breed of genomics studies designed to understand the implications of our ability to read the language of life.
Reading a genome sequence is like consulting an oracle in many ways—the messages must be interpreted carefully and often in the absence of a clear, or even complete, message. What actions should we take based on the word of the genomic oracle? How sure can we be? How much information can we currently glean? In the short term: perhaps not too much.
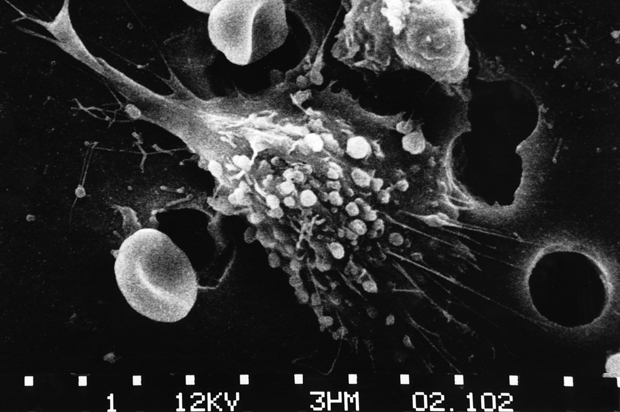
Some genes speak loud and clear, most don’t—certainly not yet given our limited knowledge of the language of DNA. The classic textbook example of a very clear message is encoded in the gene that causes the disease sickle cell anaemia. Sickle cell anaemia is caused by a single amino acid mutation in the haemoglobin gene found on chromosome 1. This causes a normally hydrophilic amino acid, glutamic acid, to be replaced with the hydrophobic amino acid, valine. The result is a sickle-shaped cell with a rigid membrane. The sickle cells are not elastic enough to allow the oxygen carrying red blood cells to flow easily through the smallest blood vessels, and this causes blockages.
There are many other genetic signatures that can have clear health implications. For instance, there is the growing pharmacogenomics database to help guide drugs better. Some 100 drugs now have a Federal Drug Administration (FDA) label recommending a genomic assessment before the drug is used.
Some genes only suggest the probability of certain outcomes. Take, for example, the ApoE gene, the strongest genetic determinant of Alzheimer’s disease and the gene that James Watson and Steven Pinker hid from themselves. Most genetic messages are strongly context-dependent. Predictions must be viewed through a maze of complex interactions of genes, environmental factors, random events, and historical contingencies. Many genes interact so heavily with external factors we can’t yet understand the consequences of possessing them. These could range from aspects of the surrounding environment, such as where you live and how much pollution you encounter, to your behaviour, such as whether you smoke or not.
If we don’t like the prophecy, how might we cope? The hope of genomic knowledge is that it will improve our lives and this includes not just mitigating any genetic shortcomings, but perhaps outright fixing them. Gene therapy is an active area of research for changing nature’s blueprint and chromosomal therapies are also being explored, for example, to cure Down’s syndrome, a genetic abnormality that causes severe learning disabilities.
The condition is caused by an extra copy of chromosome 21. In 2013, it was shown that it was possible to shut down the extra chromosome in cells from individuals with Down’s syndrome grown in the lab. This was accomplished by inserting the XIST gene, which usually inactivates one of the X chromosomes in women, into the unwanted chromosome. Now that we are getting more adept at reading genomes, we are beginning to recognize what underlies certain health outcomes. Like plastic surgery for the body, will we be able to sculpt our DNA one day?
Devil’s Ark
There is one version of our genome we hope never to see: a sad, dismantled, disgruntled, and selfish genome. Finding a cure for human cancer is the driving force in genomics. Nowhere is more effort going into understanding the process of genomic change than in human cancers.
Each cancer has a different aetiology, or origin, and a different progression of disease and expected outcomes. There are hundreds of defined cancers and tumour types because cancer can stem from any type of cells; humans possess at least 200 different types. The International Cancer Genome Consortium (ICGC), an intergovernmental collaboration, is generating a reference collection of genomes from different types of cancers, tissues, and individuals. Even this billion dollar project is only the tip of the iceberg.
Cancers are so pernicious because they achieve immortality—they don’t die but acquire superhuman powers. Due to this fact, the most famous genome in history is the HeLa genome. HeLa cell cultures have been used to underpin research that has contributed to more than 60,000 scientific articles. Bought and sold, they underpin a huge industry. They helped to develop vaccines like the one for polio, cancer medications, in vitro fertilization, gene mapping, and cloning. A year before Yuri Gagarin became the first man to orbit the Earth in 1961, HeLa cells were sent into space by the Soviets to explore how tissues respond to zero gravity.
HeLa cells have a long and contentious history. The way the HeLa genome became immortal demonstrated the tension that can arise between personal privacy and the public benefit derived through biomedical research. HeLa cells are named after Henrietta Lacks, a young woman from Virginia with five children. After a routine biopsy of her exceptionally aggressive cervical cancer cells was taken in 1951, they were walked down the hall and put into experimental culture. They proved the first to live for any significant amount of time; when cells are healthy they quickly die.
Despite the incredible importance of her cells to science, the Lacks family were kept in the dark about the existence and use of HeLa cell cultures until 25 years after her death. They were finally told only because researchers needed family DNA to create diagnostic markers for HeLa cells lines. Scientists were worried because HeLa cells were acting inexplicably. Other independently derived laboratory cell cultures, ranging from breast and prostate cells lines, turned out to be in fact HeLa cells. HeLa cells had taken them over—they were so hardy they could float on dust particles in the air and travel on unwashed hands.
HeLa cells have a genome so different from a healthy human genome that some biologists, such as Leigh Van Valen of the University of Chicago, wanted to give them a new species name, Helacyton gartleri, even though they are derived directly from human cells. While most scientists find this extreme and unhelpful, some researchers liken cancerous cells to microbial species: unicellular organisms on their own evolutionary trajectory.
Given how important these cells are to science it was inevitable that the genome should be deciphered. In 2013, Lars Steinmetz at the European Molecular Biology Laboratory in Heidelberg, Germany published a reference version of the HeLa genome. The genome sequence provided the most detailed look to date at the genetic chaos that characterizes these cells, and to a lesser extent all cancers. HeLa genomes are chock full of extra copies of her chromosomes and riddled with errors. Large segments of DNA are shuffled, rearranging the normal order of the genes in healthy humans. Around 2,000 genes are expressed at levels higher than those of normal human tissues.
Publication of the HeLa genome proved an ethical lightning rod. Several scientists, family members, and privacy campaigners objected vehemently. The genome had been sequenced and published without getting any formal consent from the Lacks family. The fact that no consent was actually required by any of the regulatory systems in place did not assuage those who felt this was an unacceptable invasion of privacy. It was bad practice and served to highlight the need for updated regulations. Steinmetz and colleagues retracted the genome.
Rebecca Skloot, author of the best-selling book on Lacks, explained why she found the genome publication morally wrong in a New York Times article. Recognizing the benefits of genome sequencing, she pointed out the urgent need to consider privacy issues: ‘No one knows what we may someday learn about Lacks’ great-grandchildren from her genome, but we know this: the view we have today of genomes is like a world map, but Google Street View is coming very soon.’
A compromise was drawn up between between scientific benefit and individual privacy. In August 2013, the National Institutes of Health (NIH) unveiled the HeLa Genome Data Use Agreement. It stipulates that a panel, including two representatives from the Lacks family, will review applications to access this genomic data and set conditions for its use. Francis Collins, head of the NIH and leader of the public Human Genome Project, made the announcement and stated: ‘We should all count Henrietta Lacks and her family as among the greatest philanthropists of our time when you consider how they contributed to the advancement of science and our health.’ HeLa cells may add to their tremendous list of accolades credit for helping to create a policy to protect genomic privacy.
Another cancer is even more extraordinary. Looney Tunes cartoons made famous an over-muscled Tasmanian Devil who spins in a tornado of fury uttering unintelligible noises and trying to eat everything in sight—most notably Bugs Bunny. In reality, Tasmanian devils are nocturnal and resemble smallish black dogs with very thick hair. The notorious devilish yowls and behaviours are just bluff, similar to a possum playing dead. The young are even cute, playful, and easily tamed.
The Tasmanian devil, Sarcophilus harrisii, suffers an infectious scourge that might actually result in its extinction—a cancer called devil facial tumour disease (DFTD). DFTD causes bloody-looking tumours on the faces of Tasmanian devils that grow so large the animals cannot feed and end up starving. The disease has a 100 per cent mortality rate. Cancerous cells are passed by bites thought to occur during feeding or mating sessions. Unlike most cancer cells, which are notoriously ‘unique’ in their exact genetic lesions, all DFTD tumours are identical to each other; the disease originated and spread from one female Tasmanian devil.
Actions are under way to save the Tasmanian devil genome. The state of Tasmania is also working to collect uninfected devils to stock a ‘Devil’s Ark’. Genomics researchers are also working hard to protect the Tasmanian devil by interpreting the genome and protecting remaining genetic diversity. One of them, Webb Miller of Penn State, also happens to be co-author of one of the most famous tools in genomics, ‘BLAST’, which compares DNA sequences. Miller is now leading a project that is helping to design a genomic conservation strategy. The obvious strategy is to protect the pool of uninfected devils, but the dream would be to cure the cancer. The holy grail for Tasmanian devils and humans alike, is to understand genomes so well we can fix them.
Excerpted from “Biocode: The New Age of Genomics” by Dawn Field & Neil Davies. Copyright © 2014 by Dawn Field & Neil Davies. Reprinted by arrangement with Oxford University Press, a division of Oxford University. All rights reserved.